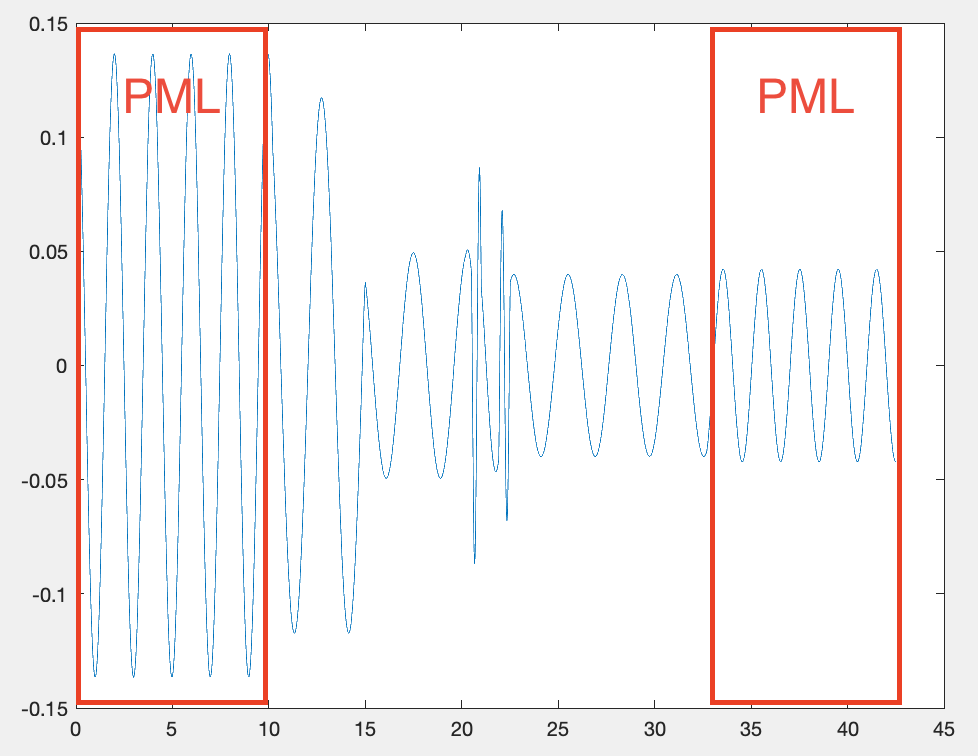
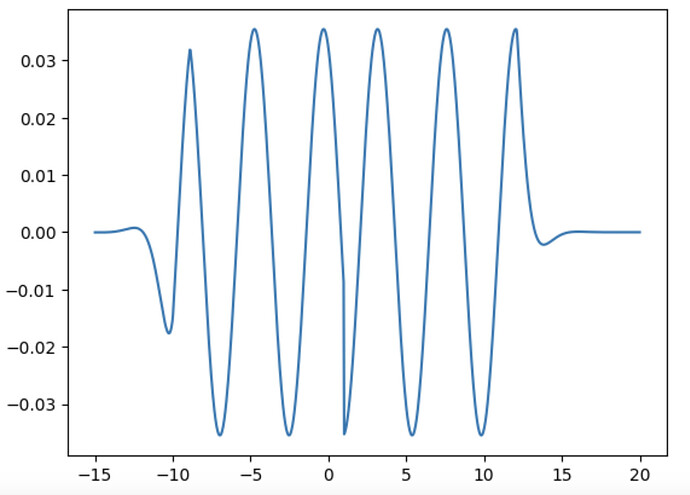
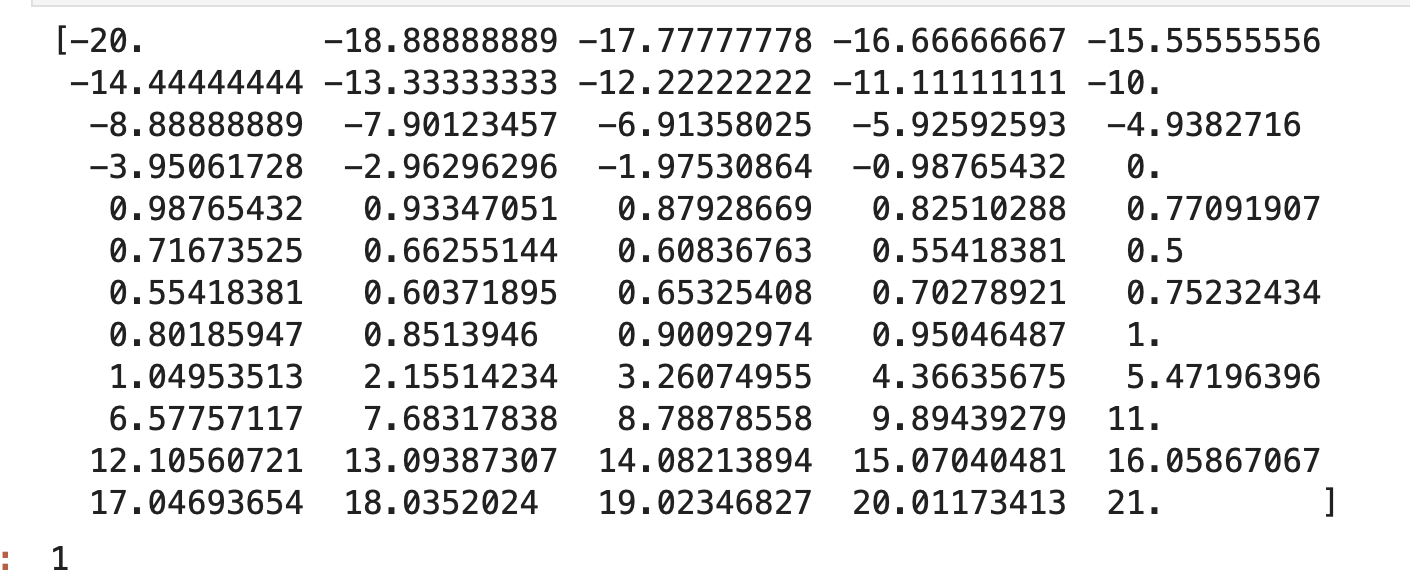
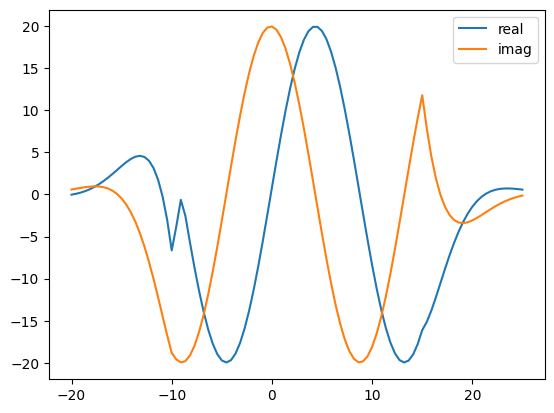
Yes. Here is the 1D wave propagation problem that I am trying to solve. Thank you.
#%%
from ngsolve import *
import ngsolve as ngs
from netgen.meshing import *
import numpy as np
import scipy.io
#%% Geometry
nnodes = 10
part = [-20, -10, 0, 0.5, 1, 11, 21]
n = []
for i in range(len(part) - 1):
n.append(nnodes)
unit_cell = Mesh(dim = 1)
diff = 0
nodes = np.empty(0)
for i in range(len(n)):
a = np.linspace(part[i] + diff, part[i + 1], n[i])
nodes = np.append(nodes, a)
diff = np.abs(nodes[len(nodes) - 1] - nodes[len(nodes) - 2])
# nodes = np.delete(nodes, 0)
pnums = []
for i in range(sum(n)):
pnums.append(unit_cell.Add(MeshPoint(Pnt(nodes[i], 0, 0))))
idx1 = unit_cell.AddRegion("pml", dim=1)
idx2 = unit_cell.AddRegion("hom", dim=1)
idx3 = unit_cell.AddRegion("rt1", dim=1)
idx4 = unit_cell.AddRegion("rt2", dim=1)
for i in range(len(nodes)-1):
if nodes[i] <= part[1]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx1))
elif nodes[i] > part[1] and nodes[i] <= part[2]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx2))
elif nodes[i] > part[2] and nodes[i] <= part[3]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx3))
elif nodes[i] > part[3] and nodes[i] <= part[4]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx4))
elif nodes[i] > part[4] and nodes[i] <= part[5]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx2))
elif nodes[i] > part[5] and nodes[i] <= part[6]:
unit_cell.Add (Element1D([pnums[i], pnums[i+1]], index = idx1))
id5 = unit_cell.AddRegion("l", dim=0)
id6 = unit_cell.AddRegion("r", dim=0)
unit_cell.Add (Element0D(pnums[0] , index=id5))
unit_cell.Add (Element0D(pnums[-1], index=id6))
#%% Meshing
mesh = ngs.Mesh(unit_cell)
mesh.SetPML(pml.Cartesian(mins = -10, alpha=1j, maxs = 11), 'pml')
#%% Materials
rhd = {'pml' : 1, 'hom' : 1, 'rt1' : 1, 'rt2' : 1}
mud = {'pml' : 2, 'hom' : 2, 'rt1' : 2, 'rt2' : 2}
rho = CoefficientFunction([rhd[mat] for mat in mesh.GetMaterials()])
mu = CoefficientFunction([mud[mat] for mat in mesh.GetMaterials()])
#%% Solution
omega = 2
fes = H1(mesh, order=3, complex=True)
u = fes.TrialFunction()
v = fes.TestFunction()
b = (10**2 / np.pi) * exp(-50**2*((x+7)**2))
a = BilinearForm(fes)
a += mu*grad(u)*grad(v)*dx - omega**2*rho*u*v*dx
f = LinearForm(fes)
f += b*v*dx # + t*v*ds("dir_neu")
a.Assemble()
f.Assemble()
gfu = GridFunction(fes)
gfu.vec.data += a.mat.Inverse(freedofs=fes.FreeDofs()) * f.vec
1D_wav_prop.py (2.7 KB)